
The Seven European Daughters
of Eve Matriarchal Groups
were given their names by Prof. Bryan Sykes:
1) Helena: This clan lived in the ice-capped Pyrenees. As the climate warmed, Helenas descendants trekked northward to what is now England, some 12,000 years ago. Members of this group are now present in all European countries.
2) Jasmine: Her people had a relatively happy life in Syria, where they farmed wheat and raised domestic animals. Jasmines descendants traveled throughout Europe, spreading their agricultural innovations with them.
3) Katrine: Members of this group lived in Venice 10,000 years ago. Today most of Katrines clan lives in the Alps.
4) Tara: Sykes maternal ancestry goes back to this group, which settled in Tuscany 17,000 years ago. Descendants ventured across northern Europe and eventually crossed the English Channel.
5) Ursula: Users of stone tools, Ursulas clan members drifted across all of Europe.
6) Valda: Originally from Spain, Valda and her immediate descendants lived 17,000 years ago. Later relatives moved into northern Finland and Norway.
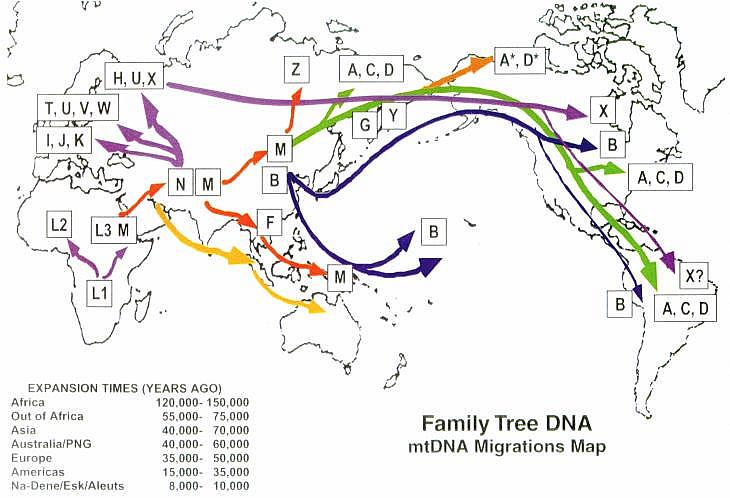
7) Xenia: Her people lived in the Caucasus Mountains 25,000 years ago. Just before the Ice Age, this clan spread across Europe, and even reached the Americas. [As Dr. Wallace discovered, the X pattern is a rare European lineage and is also among the Northern Native Americans such as the Ojibwa and Sioux.]
| Haplogroup: |
%
of Europeans: |
Years
Before
Present: |
Origin: |
| H
= Helena |
47 |
20,000 |
Southern
France |
| J
= Jasmine |
17 |
10,000 |
Middle
East |
| K
= Katrine |
6 |
15,000 |
Northern
Italy |
| T
= Tara |
9 |
17,000 |
Tuscany |
| U
= Ursula |
11 |
45,000 |
Greece |
| V
= Valda |
5 |
17,000 |
Northern
Spain |
| X
= Xenia |
6 |
25,000 |
Georgia
-
Asia |
| I
= Ina (new - 8th) |
<2 |
26,000 |
Unknown |
| CRS
Letter Reference: Base Pairs |
|
| A |
Adenine |
| C |
Cytosine |
| G |
Guanine |
| T |
Thymine |
| Mitochondrial
DNA
Results for Kit Number: |
|||||||||
| ~ HVR1 ~ (Low Resolution Haplogroup and Mutations) |
|||||||||
| Cambridge
Reference
Sequence (CRS) in Small Letters.
Participant's Sequence in Capitols. |
|||||||||
| 16010 |
16020 |
16030 |
16040 |
16050 |
16060 |
16070 |
16080 |
16090 |
16100 |
|
attctaattt |
aaactattct |
ctgttctttc |
atggggaagc |
agatttgggt |
accacccaag |
tattgactca |
cccatcaaca |
accgctatgt |
atttcgtaca |
| 16110 |
16120 |
16130 |
16140 |
16150 |
16160 |
16170 |
16180 |
16190 |
16200 |
| ttactgccag |
ccaccatgaa |
tattgtacgg |
taccataaat |
acttgaccac |
ctgtagtaca |
taaaaaccca |
atccacatca |
aaaccccctc |
cccatgctta |
| 16210 |
16220 |
16230 |
16240 |
16250 |
16260 |
16270 |
16280 |
16290 |
16300 |
| caagcaagta |
cagcaatcaa |
ccctcaacta |
tcacacatca |
actgcaactc |
caaagccacc |
cctcacccac |
taggatacca |
acaaacctac |
ccacccttaa |
| 16310 |
16320 |
16330 |
16340 |
16350 |
16360 |
16370 |
16380 |
16390 |
16400 |
| cagtacatag |
tacataaagc |
catttaccgt |
acatagcaca |
ttacagtcaa |
atcccttctc |
gtccccatgg |
atgacccccc |
tcagataggg |
gtcccttgac |
| 16410 |
16420 |
16430 |
16440 |
16450 |
16460 |
16470 |
16480 |
16490 |
16500 |
| caccatcctc |
cgtgaaatca |
atatcccgca |
caagagtgct |
actctcctcg |
ctccgggccc |
ataacacttg |
ggggtagcta |
aagtgaactg |
tatccgacat |
| 16510 |
16520 |
16530 |
16540 |
||||||
| ctggttccta |
cttcagggtc |
ataaagccta |
aatagcccac |
||||||
| ~ HVR2 ~ (High Resolution Haplogroup and Mutations) |
|||||||||
| Cambridge Reference
Sequence (CRS) in Small Letters.
Participant's Sequence in Capitols. |
|||||||||
| 00070 |
00080 |
00090 |
00100 |
00110 |
00120 |
00130 |
00140 |
00150 |
00160 |
| cgtctggggg |
gtatgcacgc |
gatagcattg |
cgagacgctg |
gagccggagc |
accctatgtc |
gcagtatctg |
tctttgattc |
ctgcctcatc |
ctattattta |
| 00170 |
00180 |
00190 |
00200 |
00210 |
00220 |
00230 |
00240 |
00250 |
00260 |
| tcgcacctac |
gttcaatatt |
acaggcgaac |
atacttacta |
aagtgtgtta |
attaattaat |
gcttgtagga |
cataataata |
acaattgaat |
gtctgcacag |
| 00270 |
00280 |
00290 |
00300 |
00310 |
00320 |
00330 |
00340 |
00350 |
00360 |
| ccactttcca |
cacagacatc |
ataacaaaaa |
atttccacca |
aaccccccct |
cccccgcttc |
tggccacagc |
acttaaacac |
atctctgcca |
aaccccaaaa |
| 00370 |
00380 |
00390 |
00400 |
00410 |
00420 |
00430 |
00440 |
00450 |
00460 |
| acaaagaacc |
ctaacaccag |
cctaaccaga |
tttcaaattt |
tatcttttgg |
cggtatgcac |
ttttaacagt |
caccccccaa |
ctaacacatt |
attttcccct |
| 00470 |
00480 |
00490 |
00500 |
00510 |
00520 |
00530 |
00540 |
00550 |
00560 |
| cccactccca |
tactactaat |
ctcatcaata |
caacccccgc |
ccatcctacc |
cagcacacac |
acaccgctgc |
taaccccata |
ccccgaacca |
accaaacccc |
| 00570 |
|||||||||
| aaagacaccc |
|||||||||
| ~ Participant Results Summary ~ | |||||||||
| Haplogroup: |
Age (Years Ago): |
%
of Europeans |
Origin: |
||||||
| Participant
# 7278 |
Participant
# 7328 |
| Participant
# 7420 |
Participant
# 8338 |